Appearance
Scatter Plot
Core Feature
This feature is available in all Portrai Explorer deployments.
The Scatter Plot extension provides interactive two-dimensional visualizations for exploring relationships between features in your data. Configure custom axes, apply color mapping, and use lasso selection directly on plots.
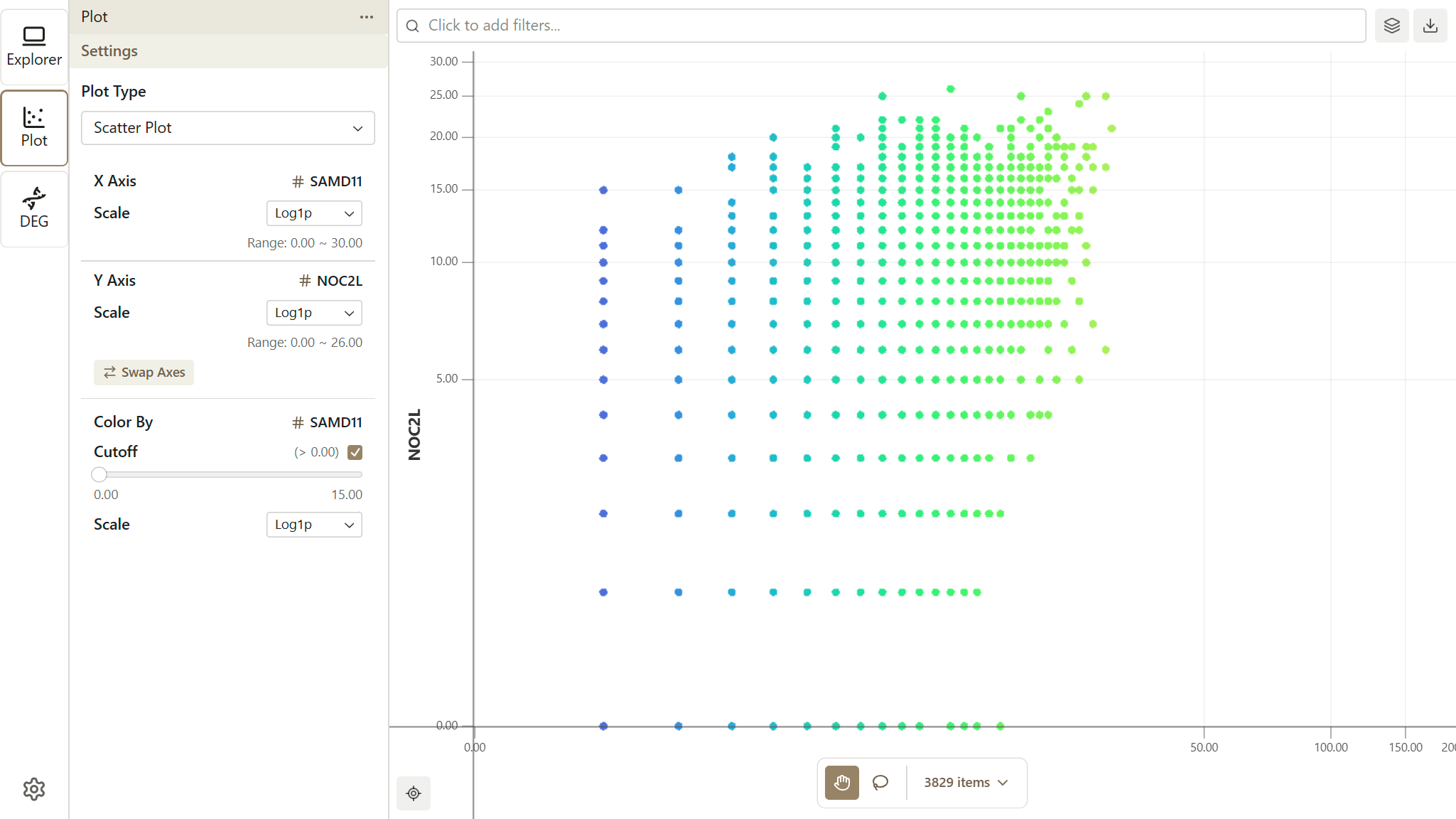
Opening the Plot Extension
- Click the Plot icon in the Activity Bar (left side)
- The Sidebar shows plot configuration options
- The Workspace displays the scatter plot
Configuring Axes
X Axis
Select what to display on the horizontal axis:
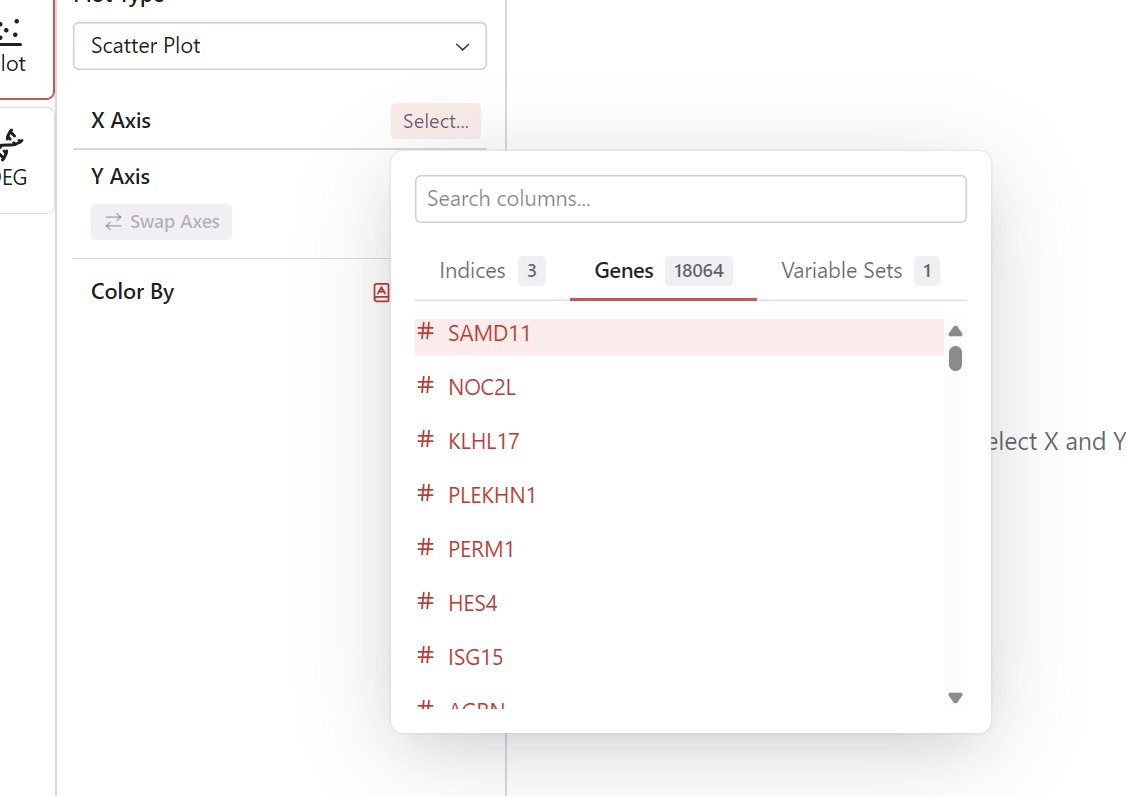
- Click the X Axis selector
- Choose from:
- Continuous features (gene expression, metrics, embedding coordinates)
- Signature scores
- The plot updates with the new X axis
TIP
Use the Sort button to organize features alphabetically when browsing long feature lists.
Y Axis
Select what to display on the vertical axis:
- Click the Y Axis selector
- Choose from the same options as X axis
- The plot updates with the new Y axis
Signature Axes
When selecting a Signature for an axis:
- Choose a Signature from the selector
- Select a Method to combine variables:
| Method | Description |
|---|---|
| Max | Maximum value among variables |
| Min | Minimum value among variables |
| Sum | Sum of variable values |
| Average | Average of variable values (default) |
| UMI Count | LogNorm: ln(10000 × sum/totalUMI + 1) |
- Configure the scale type as needed
Swap Axes
Click the Swap Axes button to exchange X and Y axis configurations, including their scale settings.
Common Axis Combinations
| X Axis | Y Axis | Purpose |
|---|---|---|
| Gene A | Gene B | Compare expression of two genes |
| Gene | n_counts | Expression vs. sequencing depth |
| Signature A | Signature B | Compare pathway scores |
Scale Types
For each axis, choose an appropriate scale:
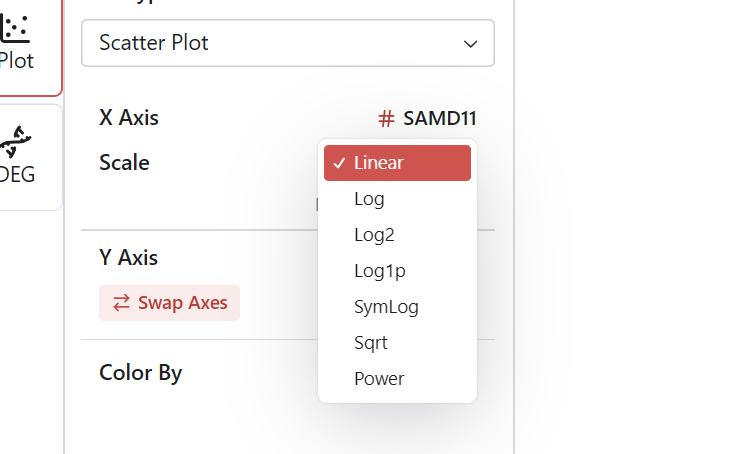
| Scale | Best For |
|---|---|
| Linear | Most data (default) |
| Log | Wide-ranging positive values |
| Log2 | Wide-ranging positive values (base 2) |
| Log1p | Data with zeros, single-cell RNA-seq |
| SymLog | Data with negative values or zeros |
| Sqrt | Moderate compression |
| Power | Emphasize differences in small values |
TIP
Log and Log2 scales are disabled when the minimum value is negative. Use SymLog or Log1p for data that includes zero or negative values.
Changing Scale
- Locate the scale selector below each axis configuration
- Select the desired scale type
- The plot redraws with the new scaling
Color Mapping
Apply color to points based on feature values:
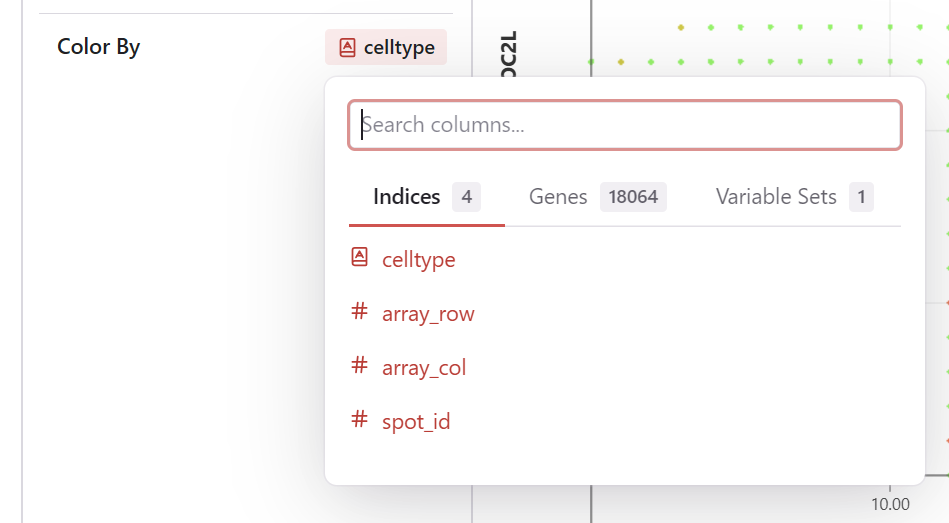
Setting Color
- Find the Color By selector in the sidebar
- Choose a feature:
- Categorical: Points colored by category
- Continuous: Points colored by value gradient
- Signature: Points colored by computed score
- The plot updates with colors
Continuous Color Settings
For continuous features and Signatures:
- Cutoff: Enable and adjust to filter out low values from the color scale (0-50%)
- Scale: Choose the scale type for color mapping
- Method (Signatures only): Select the combination method
Legend
The legend shows:
- Category names and colors (categorical)
- Color gradient with value range (continuous)
Plot Settings
Access plot settings through the Layers button (layer icon) in the top-right toolbar.
Feature Layer
Controls the opacity of all data points:
| Control | Description |
|---|---|
| Opacity slider | 0 (transparent) to 1 (opaque) |
| Eye toggle | Quick show/hide toggle |
Point Size
| Control | Description |
|---|---|
| Auto toggle | Enable automatic point sizing |
| Size slider | Manual size from 1px to 15px |
Defaults:
- Auto: Enabled
- Size: 3px (when Auto is disabled)
TIP
Manually adjusting the size slider will automatically disable Auto mode.
Axis Title Size
Adjust the font size of axis titles (10-24px, default 14px).
Details Panel
The Plot extension includes a Details Panel in the Sidebar that shows information about the current color-by selection:
- Feature color-by: Displays feature statistics (category counts for categorical, value range for continuous)
- Signature color-by: Displays Signature details including variable list, match status, and combination method controls
The Details Panel is always visible and updates automatically when you change the color-by setting.
Interacting with the Plot
Navigation
| Action | Control |
|---|---|
| Pan | Click and drag |
| Zoom | Scroll wheel |
| Reset | Click the reset button (bottom-left) |
TIP
Hold the Space key to temporarily switch to Pan mode while in Selection mode.
Hover Information
Hover over points to see:
- Point identifier (Spot ID)
- X axis column name and value
- Y axis column name and value
- Color feature name and value
- Individual variable values (if coloring by Signature)
Lasso Selection on Scatter Plots
Select points directly on the scatter plot using lasso selection.
How to Select
- Click the Selection button (Lasso icon) in the mode switcher at bottom center
- Click and drag to draw a closed shape
- Release to complete selection
- Selected points are highlighted
Using Plot Selections
Selections made on scatter plots can be:
- Added to the Subset as lasso conditions
- Used to count specific populations
- Combined with other subset conditions
Selection Context
Lasso selections on scatter plots include:
- X axis column name
- Y axis column name
- X and Y scale types
- Polygon coordinates
This information is saved so the selection can be accurately reapplied.
Export
Export the scatter plot as an image:
- Click the Export button (download icon) in the top-right toolbar
- Click Capture PNG
- The image is downloaded with axis labels and legend included
The exported image includes:
- All visible data points with current coloring
- Axis lines, labels, and titles
- Color legend (if color mapping is active)
Use Cases
Gene-Gene Comparison
Compare expression of two genes:
- Set X = Gene A expression
- Set Y = Gene B expression
- Color by cell type
- Identify co-expression patterns
Quality Control
Check data quality:
- Set X = n_counts (total counts)
- Set Y = n_genes (genes detected)
- Color by a metric or cluster
- Identify outliers or quality issues
Pathway Comparison
Compare signature scores:
- Create two Signatures
- Set X = Signature A score
- Set Y = Signature B score
- Color by cell type
- Find cells active in both pathways
Tips for Effective Plots
Choosing Axes
- Start with embeddings for overview
- Use gene pairs for co-expression analysis
- Try Signatures for pathway-level views
Optimizing Visibility
- Reduce point size for large datasets
- Lower feature layer opacity to see density
- Use log scale for expression data
Finding Patterns
- Try different color mappings
- Switch scales to reveal structure
- Use lasso to isolate clusters
Troubleshooting
Plot is Empty
- Check that X and Y axes are selected
- Verify data is loaded
- Check if subsets are excluding all points
Points Overlap Too Much
- Reduce point size
- Lower feature layer opacity
- Try different scales
Colors Not Visible
- Check ColorBy selection
- Verify feature has variation
- Adjust color range or cutoff
Slow Performance
- Large datasets may render slowly
- Try subsetting to reduce point count
- Allow time for initial render
Related Topics
- Color Mapping - Color configuration details
- Signatures - Create axis variables
- Lasso Selection - Selection techniques
- Layer Settings - Point appearance settings